
东北大学学报(自然科学版) ›› 2026, Vol. 47 ›› Issue (1): 31-41.DOI: 10.12068/j.issn.1005-3026.2026.20259019
收稿日期:2025-09-14
出版日期:2026-01-15
发布日期:2026-03-17
通讯作者:
班晓娟
作者简介:樊捷杰(1985—),男,江西余干人,北京科技大学博士研究生.
基金资助:
Jie-jie FAN1, Xiao-juan BAN1( ), Zhi-yan ZHANG2
), Zhi-yan ZHANG2
Received:2025-09-14
Online:2026-01-15
Published:2026-03-17
Contact:
Xiao-juan BAN
摘要:
现有的电子健康记录(electronic health records, EHR)的图表示学习方法多依赖单个患者的局部信息,忽视了群体患者在疾病演化和诊疗路径上的潜在关联,从而限制了模型的泛化性与鲁棒性.针对这一问题,本文提出一种混合多层级图神经网络(hybrid multi-level graph neural network, H-MGNN)模型,并将其应用于重症监护室(intensive care unit, ICU)患者的死亡预测.该模型通过构建宏观层面的患者关系图(patient-patient graph, P-P)、微观层面的分类-笔记-词汇超图(taxonomy-note-word hypergraph, T-N-W),结合超图的时序依赖关系,实现多尺度上的患者特征融合.同时,本文设计了融合算法(hybrid embedding, Hybrid-E),用于提取和整合患者嵌入的潜在特征,以提升预测准确性.实验结果表明,H-MGNN在MIMIC-Ⅲ(medical information mart for intensive care Ⅲ)数据集上的住院死亡率预测等任务中显著优于现有方法,验证了其在复杂EHR数据挖掘中的有效性和先进性.
中图分类号:
樊捷杰, 班晓娟, 张志研. 一种基于电子健康记录的多尺度图表示学习模型[J]. 东北大学学报(自然科学版), 2026, 47(1): 31-41.
Jie-jie FAN, Xiao-juan BAN, Zhi-yan ZHANG. A Multi-scale Graph Representation Learning Model Based on Electronic Health Records[J]. Journal of Northeastern University(Natural Science), 2026, 47(1): 31-41.
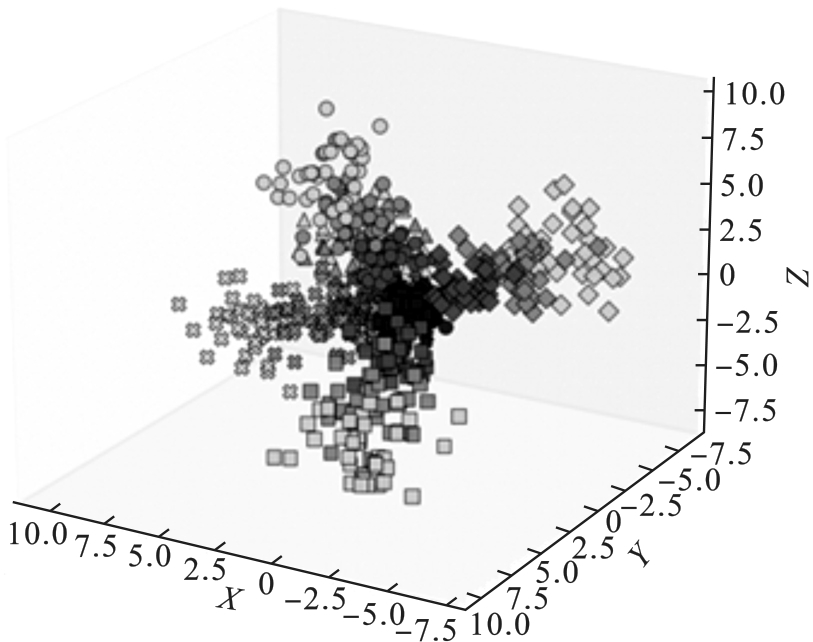
图2 患者样本3D示例(不同形状代表不同疾病,颜色深浅代表疾病的严重程度)
Fig.2 3D visualization of patient samples (different shapes represent different diseases, color intensity indicates disease severity)
| 类别 | 模型 | 整体 | 高血压 | 糖尿病 | |||
|---|---|---|---|---|---|---|---|
| AUPRC | AUROC | AUPRC | AUROC | AUPRC | AUROC | ||
| 字符 | FastText | 17.06±0.08 | 62.37±0.11 | 25.56±0.28 | 62.39±0.18 | 31.33±0.33 | 67.89±0.20 |
| 时序 | Bi-LSTM | 17.67±4.19 | 58.75±5.78 | 21.75±5.25 | 57.39±6.11 | 27.52±7.57 | 61.86±8.38 |
| Bi-LSTM w/Att | 17.96±0.61 | 62.63±1.31 | 26.05±1.80 | 63.24±1.57 | 33.01±3.53 | 68.89±1.58 | |
| 图 | TextING | 34.50±7.79 | 78.20±4.27 | 36.63±8.30 | 80.12±4.05 | 36.13±8.66 | 80.28±3.84 |
| InducT-GCN | 43.03±1.96 | 82.23±0.72 | 41.06±2.95 | 85.56±1.24 | 40.59±3.07 | 84.42±1.45 | |
| 超图 | HyperGAT | 44.42±1.96 | 84.00±0.84 | 42.32±1.78 | 86.41±1.01 | 40.08±2.45 | 85.03±1.20 |
| TM-HGNN* | 46.15±1.43 | 84.45±0.62 | 44.50±1.66 | 87.15±0.34 | 41.24±1.72 | 85.68±1.05 | |
| 本文算法 | H-MGNN | 49.80±0.54 | 86.35±0.30 | 48.67±0.72 | 87.95±0.45 | 44.62±0.80 | 87.85±1.16 |
表1 患者院内死亡率预测结果
Table 1 Prediction results of in-hospital mortality for patients
| 类别 | 模型 | 整体 | 高血压 | 糖尿病 | |||
|---|---|---|---|---|---|---|---|
| AUPRC | AUROC | AUPRC | AUROC | AUPRC | AUROC | ||
| 字符 | FastText | 17.06±0.08 | 62.37±0.11 | 25.56±0.28 | 62.39±0.18 | 31.33±0.33 | 67.89±0.20 |
| 时序 | Bi-LSTM | 17.67±4.19 | 58.75±5.78 | 21.75±5.25 | 57.39±6.11 | 27.52±7.57 | 61.86±8.38 |
| Bi-LSTM w/Att | 17.96±0.61 | 62.63±1.31 | 26.05±1.80 | 63.24±1.57 | 33.01±3.53 | 68.89±1.58 | |
| 图 | TextING | 34.50±7.79 | 78.20±4.27 | 36.63±8.30 | 80.12±4.05 | 36.13±8.66 | 80.28±3.84 |
| InducT-GCN | 43.03±1.96 | 82.23±0.72 | 41.06±2.95 | 85.56±1.24 | 40.59±3.07 | 84.42±1.45 | |
| 超图 | HyperGAT | 44.42±1.96 | 84.00±0.84 | 42.32±1.78 | 86.41±1.01 | 40.08±2.45 | 85.03±1.20 |
| TM-HGNN* | 46.15±1.43 | 84.45±0.62 | 44.50±1.66 | 87.15±0.34 | 41.24±1.72 | 85.68±1.05 | |
| 本文算法 | H-MGNN | 49.80±0.54 | 86.35±0.30 | 48.67±0.72 | 87.95±0.45 | 44.62±0.80 | 87.85±1.16 |
| 消融操作 | 模型 | 整体 | 高血压 | ||
|---|---|---|---|---|---|
| AUPRC | AUROC | AUPRC | AUROC | ||
| 去除P-P模块 | T-MGNN | 46.27±1.05 | 84.90±0.84 | 44.65±0.60 | 87.45±0.67 |
| 去除P-P及时序 | T-N-W | 45.63±1.26 | 84.23±0.55 | 43.36±0.85 | 86.68±0.78 |
| 增加内部位置 | TM-HGNN* | 46.15±1.43 | 84.45±0.62 | 44.50±1.66 | 87.15±0.34 |
| 本文算法 | H-MGNN | 49.80±0.54 | 86.35±0.30 | 48.67±0.72 | 87.95±0.45 |
表2 模块有效性消融实验结果
Table 2 Ablation experiment results of module effectiveness
| 消融操作 | 模型 | 整体 | 高血压 | ||
|---|---|---|---|---|---|
| AUPRC | AUROC | AUPRC | AUROC | ||
| 去除P-P模块 | T-MGNN | 46.27±1.05 | 84.90±0.84 | 44.65±0.60 | 87.45±0.67 |
| 去除P-P及时序 | T-N-W | 45.63±1.26 | 84.23±0.55 | 43.36±0.85 | 86.68±0.78 |
| 增加内部位置 | TM-HGNN* | 46.15±1.43 | 84.45±0.62 | 44.50±1.66 | 87.15±0.34 |
| 本文算法 | H-MGNN | 49.80±0.54 | 86.35±0.30 | 48.67±0.72 | 87.95±0.45 |
| [1] | Johnson A E W, Pollard T J, Shen L, et al. MIMIC-III, a freely accessible critical care database[J]. Scientific Data, 2016, 3: 160035. |
| [2] | Lipton Z C, Kale D C, Elkan C, et al. Learning to diagnose with LSTM recurrent neural networks[C]// International Conference on Learning Representations (ICLR). San Juan, 2016:1-8. |
| [3] | Che Z P, Purushotham S, Cho K, et al. Recurrent neural networks for multivariate time series with missing values[J]. Scientific Reports, 2018, 8: 6085. |
| [4] | Malone B, Garcia-Duran A, Niepert M. Learning representations of missing data for predicting patient outcomes[EB/OL]. (2018-12-12)[2025-02-18].. |
| [5] | Xu Y B, Biswal S, Deshpande S R, et al. RAIM: recurrent attentive and intensive model of multimodal patient monitoring data[C]//Proceedings of the 24th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining (KDD ’18). London, 2018: 2565-2573. |
| [6] | Ngo Q H, Kechadi T, Le-Khac N A. Domain specific entity recognition with semantic-based deep learning approach[J]. IEEE Access, 2021, 9: 152892-152902. |
| [7] | Rasmy L, Xiang Y, Xie Z Q, et al. Med-BERT: pretrained contextualized embeddings on large-scale structured electronic health records for disease prediction[J]. NPJ Digital Medicine, 2021, 4: 86. |
| [8] | Lee J, Yoon W, Kim S, et al. BioBERT: a pre-trained biomedical language representation model for biomedical text mining[J]. Bioinformatics, 2020, 36(4): 1234-1240. |
| [9] | Alsentzer E, Murphy J R, Boag W, et al. Publicly available clinical BERT embeddings[EB/OL]. (2019-04-06)[2025-01-10]. . |
| [10] | Vaswani A, Shazeer N, Parmar N, et al. Attention is all you need[J]. Advances in Neural Information Processing Systems, 2017, 30:6000-6010. |
| [11] | Huang K X, Altosaar J, Ranganath R. Clinical BERT: modeling clinical notes and predicting hospital readmission[EB/OL]. (2019-04-10)[2021-04-15]. . |
| [12] | Lewis P, Perez E, Piktus A, et al. Retrieval-augmented generation for knowledge-intensive NLP tasks[J]. Advances in Neural Information Processing Systems, 2020, 33: 9459-9474. |
| [13] | Song H, Rajan D, Thiagarajan J, et al. Attend and diagnose: clinical time series analysis using attention models[C]//AAAI Conference on Artificial Intelligence. New Orleans: AAAI Press, 2018: 4091-4098. |
| [14] | Hirszowicz O, Aran D. ICU bloodstream infection prediction: a transformer-based approach for EHR analysis[C]//Artificial Intelligence in Medicine. Cham: Springer, 2024: 279-292. |
| [15] | Li Y K, Rao S, Solares J R A, et al. BEHRT: transformer for electronic health records[J]. Scientific Reports, 2020, 10: 7155. |
| [16] | Pennington J, Socher R, Manning C. GLOVE: global vectors for word representation[C]// Proceedings of the 2014 Conference on Empirical Methods in Natural Language Processing (EMNLP). Doha: ACL, 2014: 1532-1543. |
| [17] | Tipirneni S, Reddy C K. Self-supervised transformer for sparse and irregularly sampled multivariate clinical time-series[J]. ACM Transactions on Knowledge Discovery from Data, 2022, 16(6): 105. 1-105.17. |
| [18] | Kipf T N, Welling M. Semi-supervised classification with graph convolutional networks[EB/OL]. (2016-09-09)[2020-01-05]. . |
| [19] | Brody S, Yahav E, Levy O. Attentive neural processes[EB/OL]. (2021-01-17)[2022-01-05]. . |
| [20] | Liu X E, You X X, Zhang X, et al. Tensor graph convolutional networks for text classification[C]// Proceedings of the AAAI Conference on Artificial Intelligence. Philadelphia: AAAI Press, 2020: 8409-8416. |
| [21] | Yao L, Mao C S, Luo Y. Graph convolutional networks for text classification[C]// Proceedings of the AAAI Conference on Artificial Intelligence. Los Angeles: AAAI Press, 2019: 7370-7377. |
| [22] | Zhang Y F, Yu X L, Cui Z Y, et al. Every document owns its structure: inductive text classification via graph neural networks[EB/OL]. (2020-04-22)[2021-05-10]. . |
| [23] | Wang K Z, Han S C, Poon J. InducT-GCN: inductive graph convolutional networks for text classification[C]// 2022 26th International Conference on Pattern Recognition (ICPR). Montreal: IEEE, 2022: 1243-1249. |
| [24] | Piao Y h, Lee S S, Lee D, et al. Sparse structure learning via graph neural networks for inductive document classification[C]//Processing of the AAAI Conference on Aritificial Intelligence.Vancouver,2022:11165-11173. |
| [25] | Ding K Z, Wang J L, Li J D, et al. Be more with less: hypergraph attention networks for inductive text classification[EB/OL]. (2020-11-01)[2023-05-10]. . |
| [26] | Zhang H P, Liu X, Zhang J W. HEGEL: hypergraph transformer for long document summarization[EB/OL]. (2022-08-09)[2023-05-10]. . |
| [27] | Park S, Bae S, Kim J, et al. Graph-text multi-modal pre-training for medical representation learning[C]// ACM Conference on Health, Inference, and Learning. Online, 2022: 261-281. |
| [28] | Zhang C H, Chu X, Ma L T, et al. M3Care: learning with missing modalities in multimodal healthcare data[C]// Proceedings of the 28th ACM SIGKDD Conference on Knowledge Discovery and Data Mining. Washington DC, 2022: 2418-2428. |
| [29] | Xu Y X, Yang K, Zhang C H, et al. VecoCare: visit sequences-clinical notes joint learning for diagnosis prediction in healthcare data[C]// Proceedings of the Thirty-Second International Joint Conference on Artificial Intelligence. Macau, 2023: 4921-4929. |
| [30] | Chen D X, O’Bray L, Borgwardt K M. Structure-aware transformer for graph representation learning[C]// International Conference on Machine Learning. Online, 2022: 3469-3489. |
| [31] | Choi E, Bahadori M T, Song L, et al. GRAM: graph-based attention model for healthcare representation learning[C]// Proceedings of the 23rd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining. Halifax NS: ACM, 2017: 787-795. |
| [32] | Qiu L, Gorantla S, Rajan V, et al. Multi-disease predictive analytics: a clinical knowledge-aware approach[J]. ACM Transactions on Management Information Systems, 2021, 12(3): 1-34. |
| [33] | Ma J T, Liu B, Li K L, et al. A review of graph neural networks and pretrained language models for knowledge graph reasoning[J]. Neurocomputing, 2024, 609: 128490. |
| [34] | Mo X, Ding G H, Tang R, et al. Bipartite graphs contrastive learning with knowledge-aware diffusion-enhanced[J]. IEEE Transaction Network Science and Engineering, 2025, 12(5): 4182-4195. |
| [35] | Mishra R, Shridevi S. Knowledge graph driven medicine recommendation system using graph neural networks on longitudinal medical records[J]. Scientific Reports, 2024, 14: 25449. |
| [36] | Gaupp R, Dinius J, Drazic I, et al. Long-term effects of an e-learning course on patient safety: a controlled longitudinal study with medical students[J]. PLoS One, 2019, 14(1): e0210947. |
| [37] | Gupta S, Sharma S, Sharma R, et al. Healing with hierarchy: hierarchical attention empowered graph neural networks for predictive analysis in medical data[J]. Artificial Intelligence in Medicine, 2025, 165: 103134. |
| [38] | Zhang D D, Yin C C, Zeng J C, et al. Combining structured and unstructured data for predictive models: a deep learning approach[J]. BMC Medical Informatics and Decision Making, 2020, 20: 280. |
| [39] | Gayathri R, Sangeetha S K B, Sangeetha R, et al. Dynamic AI-enhanced therapeutic framework for precision medicine using multi-modal data and patient-centric reinforcement learning[J]. IEEE Access, 2025, 13: 77709-77733. |
| [40] | Huang K X, Singh A, Chen S T, et al. Clinical XLNet: modeling sequential clinical notes and predicting prolonged mechanical ventilation[EB/OL]. (2019-12-27)[2020-10-10]. . |
| [41] | Hou L X, Zhuang Y, Xie Y H, et al. Cross-modal generalizable visual-language models via inter-modal bidirectional supervision for enhanced pathology image recognition[J]. Pattern Recognition, 2026, 171: 112240. |
| [42] | Hastuti R P, Rajagede R A, Zheng M, et al. Clinic-prompt: few-shot discrete clinical prompt optimization[C]//Workshop on Large Language Models and Generative AI for Health at AAAI 2025. Philadelphia, 2025:2451490. |
| [43] | Mulyar A, Schumacher E, Rouhizadeh M, et al. Phenotyping of clinical notes with improved document classification models using contextualized neural language models[EB/OL]. (2019-10-30)[2021-01-02]. . |
| [44] | Kruskal J B. Multidimensional scaling by optimizing goodness of fit to a nonmetric hypothesis[J]. Psychometrika, 1964, 29(1): 1-27. |
| [45] | Haugen E, Firth J R. Papers in linguistics 1934—1951[J]. Language, 1958, 34(4): 498-502. |
| [46] | Tenenbaum J B, de Silva V, Langford J C. A global geometric framework for nonlinear dimensionality reduction[J]. Science, 2000, 290(5500): 2319-2323. |
| [47] | Roweis S T, Saul L K. Nonlinear dimensionality reduction by locally linear embedding[J]. Science, 2000, 290(5500): 2323-2326. |
| [48] | Belkin M, Niyogi P. Laplacian eigenmaps and spectral techniques for embedding and clustering[C]// Advances in Neural Information Processing Systems 14. Cambridge, MA: MIT Press, 2002: 585-592. |
| [49] | Cao S S, Lu W, Xu Q K. GraRep: learning graph representations with global structural information[C]//Proceedings of the 24th ACM International on Conference on Information and Knowledge Management. Melbourne, 2015: 891-900. |
| [50] | Ou M D, Cui P, Pei J, et al. Asymmetric transitivity preserving graph embedding[C]//Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining. San Francisco: ACM, 2016: 1105-1114. |
| [51] | Perozzi B, Al-Rfou R, Skiena S. DeepWalk: online learning of social representations[C]//Proceedings of the 20th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining. New York: ACM, 2014: 701-710. |
| [52] | Tang J, Qu M, Wang M Z, et al. LINE: large-scale information network embedding[EB/OL]. (2015-03-12)[2020-03-11]. . |
| [53] | Tang J, Qu M, Mei Q Z. PTE: predictive text embedding through large-scale heterogeneous text networks[C]//Proceedings of the 21st ACM SIGKDD International Conference on Knowledge Discovery and Data Mining. Sydney, NSW: ACM, 2015: 1165-1174. |
| [54] | Harutyunyan H, Khachatrian H, Kale D C, et al. Multitask learning and benchmarking with clinical time series data[J]. Scientific Data, 2019, 6: 96. |
| [55] | Kim N, Piao Y H, Kim S. Clinical note owns its hierarchy: multi-level hypergraph neural networks for patient-level representation learning[EB/OL]. (2023-05-16)[2025-02-20]. . |
| [56] | Zhou P, Shi W, Tian J, et al. Attention-based bidirectional long short-term memory networks for relation classification[C]//Proceedings of the 54th Annual Meeting of the Association for Computational Linguistics. Berlin: ACL, 2016: 207-212. |
| [57] | Wang Z H, Yang B. Attention-based bidirectional long short-term memory networks for relation classification using knowledge distillation from BERT[C]// 2020 IEEE International Conference on Dependable, Autonomic and Secure Computing, International Conference on Pervasive Intelligence and Computing, international conference on Cloud and Big Data Computing, international conference on Cyber Science and Technology Congress (DASC/PiCom/CBDCom/CyberSciTech). Calgary: IEEE, 2020: 562-568. |
| [58] | Joulin A, Grave E, Bojanowski P, et al. Bag of tricks for efficient text classification[EB/OL]. (2016-07-06)[2023-05-10]. . |
| [1] | 李立振, 马淑华, 郭泽旭, 车晓辰. 基于X-ray-RTDETR的X射线图像违禁品检测算法[J]. 东北大学学报(自然科学版), 2025, 46(6): 8-15. |
| [2] | 李鸿儒, 李同同, 石康康, 杨英华. 基于HRV多尺度分析的糖尿病前期检测方法[J]. 东北大学学报(自然科学版), 2025, 46(12): 19-28. |
| [3] | 冯虎, 宋克臣, 崔文琦, 颜云辉. 基于元学习的带钢表面缺陷小样本语义分割[J]. 东北大学学报(自然科学版), 2024, 45(3): 354-360. |
| [4] | 单鹏, 张林, 肖洪明, 赵玉良. 融合多尺度注意力机制的冠状病毒肺炎CT诊断方法[J]. 东北大学学报(自然科学版), 2024, 45(12): 1673-1679. |
| [5] | 冯昂, 宫俊, 王念, 王景龙. 基于图卷积和卷积的行人轨迹预测算法[J]. 东北大学学报(自然科学版), 2024, 45(11): 1529-1536. |
| [6] | 吕艳霞, 郝帅, 乔广通, 邢烨. 一种基于图神经网络的社会化推荐算法[J]. 东北大学学报(自然科学版), 2024, 45(1): 10-17. |
| [7] | 李海燕, 熊立昌, 郭磊, 李海江. 基于U-net边缘生成和超图卷积的两阶段修复算法[J]. 东北大学学报(自然科学版), 2023, 44(3): 331-339. |
| [8] | 金瑾, 王鹏. 历史数据驱动的多尺度量子谐振子优化算法[J]. 东北大学学报(自然科学版), 2022, 43(2): 160-167. |
| [9] | 连静, 陈实, 丁堃, 李琳辉. 基于多尺度密集特征融合的生成式对抗除雾网络[J]. 东北大学学报(自然科学版), 2022, 43(11): 1591-1598. |
| [10] | 张雪峰, 何昊. 基于分数阶微分的多尺度多聚焦图像融合[J]. 东北大学学报(自然科学版), 2021, 42(8): 1071-1078. |
| [11] | 原培新, 陈鼎夫. 双能X射线高动态范围安检图像压缩算法[J]. 东北大学学报(自然科学版), 2021, 42(1): 96-101. |
| [12] | 杨丹, 刘国如, 任梦成, 裴宏杨. 多尺度卷积核U-Net模型的视网膜血管分割方法[J]. 东北大学学报(自然科学版), 2021, 42(1): 7-14. |
| [13] | 张云洲, 郑瑞, 暴吉宁, 朱尚栋. 基于多个相关滤波器的行人跟踪尺度算法[J]. 东北大学学报:自然科学版, 2019, 40(9): 1228-1234. |
| [14] | 龙哲, 王旭, 杨丹. 交变磁场曝露对大鼠心电信号多尺度熵的影响[J]. 东北大学学报:自然科学版, 2019, 40(6): 766-770. |
| [15] | 刘威, 遇冰, 周婷, 袁淮. 基于多特征融合的图像区域几何标记[J]. 东北大学学报:自然科学版, 2017, 38(7): 927-931. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||